


Notice
PDB news
Sussman New Director of PDB
Joel L. Sussman of the Weizmann Institute of Science in Israel is the newly appointed head of the PDB. Joel was one of the early developers of methodology and programs on biological macromolecular refinement. Recently he has represented the European Science Foundation Network on the PDB Advisory Board. Sussmann envisions PDB as a true international collaboration, receiving not only data, but a free flow of ideas from its growing community of users. Tom Koetzle, the former Director of the PDB will continue as a consultant to the new PDB management.
 Database gurus Frank Allen (CSD) and Tom Koetzle (PDB) compare notes in Beijing, 1993. (Photo WLD)
Database gurus Frank Allen (CSD) and Tom Koetzle (PDB) compare notes in Beijing, 1993. (Photo WLD)
The 605 new additions to the January 1994 PDB Release brought the total number of full-release atomic coordinate entries to 2327. These included 2143 proteins, enzymes, and viruses, 156 DNA's, 9 RNA's, 9 tRNA's, and 10 carbohydrates. The total size of the atomic coordinate entry database is 678 Mbytes. New software has been installed by PDB to assist with data processing and checking, to automate a variety of processing tasks and tp improve quality control, ensuring stricter adherence to PDB's published specifications.
Significant gains in data processing productivity have been achieved, enabling PDB to respond to the challenge posed by the exponentially growing number of structure submissions. The staff of the PDB are actively soliciting software from the user community. If you have data preparation or checking programs that you are willing to share, please contact PDB directly at the address given below.
Windows Users
PDB is pleased to announce a new facility for Windows users to access and display structures from the PDB. This new facility is called PDB-Shell and is available on the January1994 CD-ROM and on the PDB anonymous FTP server.
CD-ROM Info.
PDB releases are available on CD-ROM mirrors in ISO 9660 format. The layout of files on the CD-ROM mirrors the UNIX tape distribution and uses the tape01, tape02, ..., tapeN subdirectories. The entry files are ASCII format and are readable by software able to read text files.
Procheck Software Package
The Procheck software package is being made available for electronic distribution from PDB. Oxford Molecular, Ltd and PDB have agreed that upon receipt of a signed license agreement at PDB, the source and documentation for Procheck will be made available free of charge. The Procheck software package, created by J. M. Thornton, M. W. MacArthur, R. A. Laskowski, and D. S. Moss, performs evaluations of the stereochemical quality of protein structures.
In order to acquire a copy of Procheck, you must obtain the license agreement, copies of which are available via the PDB FTP, gopher, or file serve in the file called procheck-license. You must complete and sign this license agreement and return it to PDB. Once they have your signed agreement, they will either e-mail the source to you or place it on the machine of your choice via FTP. Your signed license agreement will be forwarded to Oxford Molecular who will keep you up to date about further developments. All queries concerning the software should be directed to Steve Gardner, Macromolecular Product Manager, Oxford Molecular, Ltd, The Magdalen Centre, Oxford Science Park, Sanford-on Thames, Oxford, England OX4 4GA. Tel.: 44-865-784600.
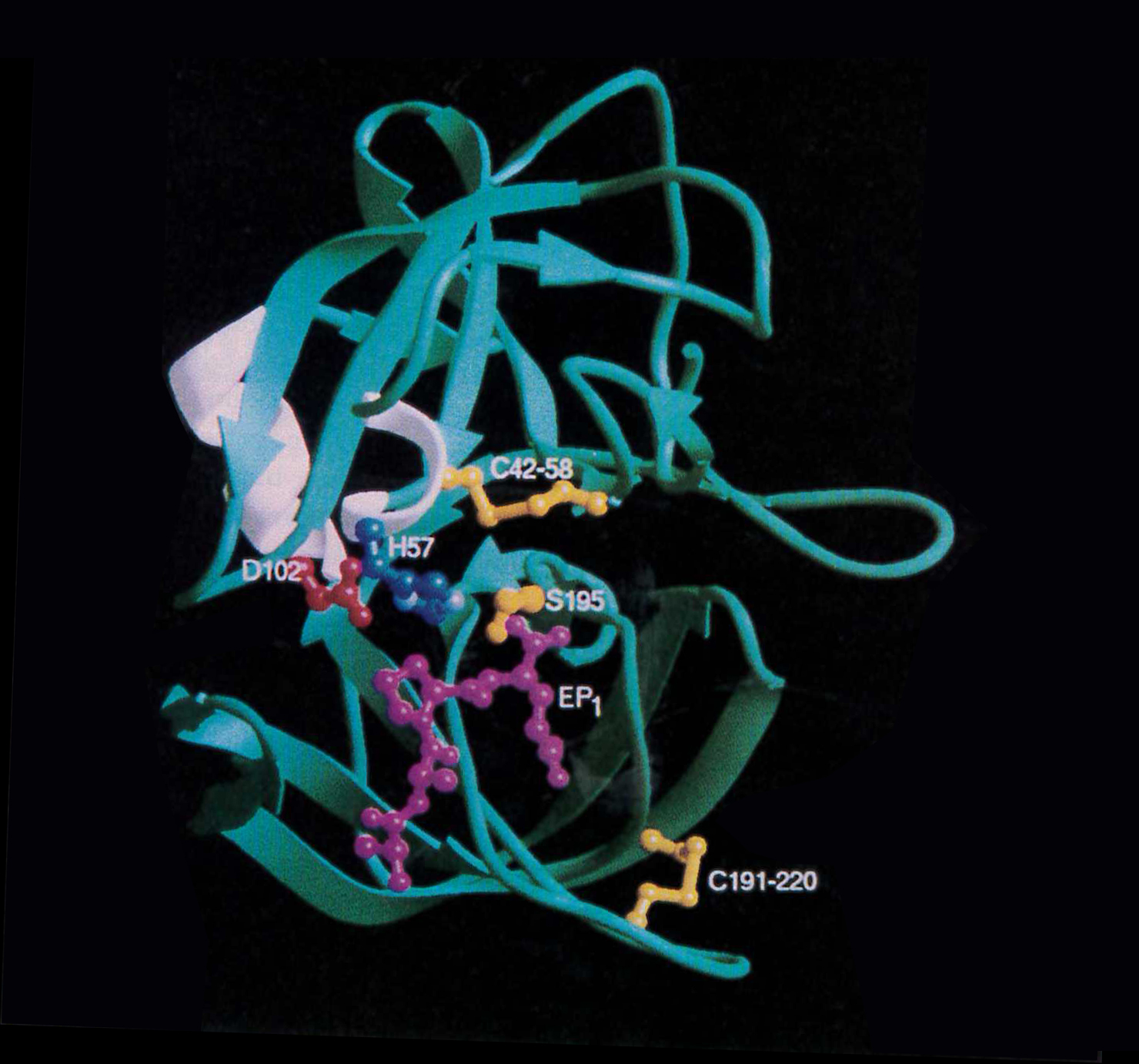 Glutamic acid specific protease from Strept griseus. Helices are shown in white, while β-strands are drawn as green arrows. The N-terminal β-barrel 1 forms the top portion of the structure while β-barrel 2 containing the substrate binding site is located at the bottom. The catalytic triad (Asp102, His57, and Ser195), color coded red, blue, and yellow, respectively, and the tetrapeptide ligand (Ala-Ala-Pro-Glu), in pink, are represented as ball and stick models. The two disulfide bridges, Cys42-Cys58 in β-barrel 1 (top) and Cys191-Cys220 in β-barrel 2 (bottom) are also shown. The atomic coordinates are on deposit in the Protein Data Bank, Brookhaven, PDB code 1HPG (from Biochemistry, 1993, 32, 11469-11475).
Glutamic acid specific protease from Strept griseus. Helices are shown in white, while β-strands are drawn as green arrows. The N-terminal β-barrel 1 forms the top portion of the structure while β-barrel 2 containing the substrate binding site is located at the bottom. The catalytic triad (Asp102, His57, and Ser195), color coded red, blue, and yellow, respectively, and the tetrapeptide ligand (Ala-Ala-Pro-Glu), in pink, are represented as ball and stick models. The two disulfide bridges, Cys42-Cys58 in β-barrel 1 (top) and Cys191-Cys220 in β-barrel 2 (bottom) are also shown. The atomic coordinates are on deposit in the Protein Data Bank, Brookhaven, PDB code 1HPG (from Biochemistry, 1993, 32, 11469-11475).
PDB via New Listserver
The Protein Data Bank has formed a mailing list devoted to discussions concerning the operation of the data bank, its contents, and access to it. If you wish tosubscribe, send e-mail to [email protected] with the message: subscribe PDB-L firstname lastname. To find out what you can do with this mailing list, send e-mail to the same address ([email protected]) with the one line message: HELP. To send a message to all PDB-L subscribers, e-mail message to [email protected].
For further information contact: Protein Data Bank, Chem. Dept., Bldg. 555, Brookhaven National Laboratory, PO Box 5000, Upton, NY 11973-5000, USA. Tel.: 516-282-3629, FAX: 516-282-5751, e-mail: [email protected] or [email protected].
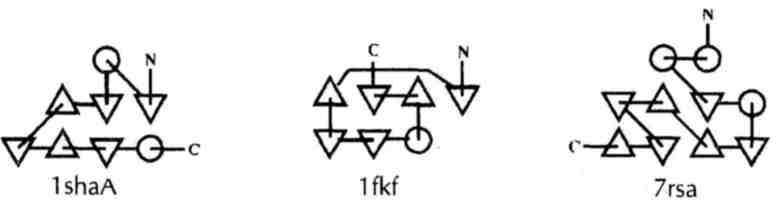
Taken from Protein Data Bank Quarterly Newsletter, October 1993 and January 1994.


